Background
Decoy and Tough Decoy (TuD) RNAs are novel technologies for regulating RNA interference (Haraguchi et al., Nucleic Acids Res. 2009; Xie et al., Nature 2012). In short, they are single strands of RNA with one antisense microRNA binding domain (Decoy) or a stabilized stem-loop with two microRNA binding domains (TuD).
In accordance with the central dogma of biology, transcribed mRNA strands are translated by cellular machinery into proteins. Mammalian cells incorporate a layer of expression control, RNA interference, to control mRNA levels. RNA interference uses microRNAs (miRNAs) that are processed by protein complexes to knock down mRNA levels in the cell, reducing protein expression. Decoys and TuDs are artificial strands of RNA with miRNA-binding domains that are thought to sequester the miRNA into stable complexes throuh complementary basepairing, disabling a particular RNA interference pathway.
While miRNAs act as repressors, TuDs and Decoys act as double-repressors such that the presence of the TuDs/Decoys increases protein output.
Thus miRNAs, Decoys and TuDs can potentially be incorporated in RNA strand-displacement circuits to control protein expression levels. As these are short RNAs, we imagine that these strands can be outputs of RNA strand displacement circuits. This will allow strand-displacement circuits to connect to traditional protein-based synthetic biology circuits.

Diagram depicting the interactions between Decoys, TuDs, miRNAs, mRNAs and proteins.
miRNAs are used in the RNA interference pathway to knock down mRNAs, which are usually translated into proteins. Decoys and TuDs repress this pathway, resulting in protein production.
Experimental Design
We designed a proof-of-concept double-repression circuit to test the in vivo actuation ability of Decoy and TuD RNAs as indicated by fluorescent protein expression.
We observed from the Haraguchi et al. study that a particular TuD RNA structure, one with a stabilizing stem-loop structure, yielded increased levels of activity compared to Decoy RNA, so we set out to assess the functionalities of both Decoy and TuD designs. Furthermore, the studies showed that a small 4-nucleotide bulge sequence in the miRNA-binding domains of Decoy and TuD RNAs increased their abilities to relieve repression of protein expression. The bulge in the miRNA-binding sequence likely protected the Decoy and TuD RNA from the RISC protein complex. This characteristic informed our decision to test two different Decoy and TuD RNA species – one that lacked a 4-nucleotide bulge in the miRNA-binding domain, and one that had the bulge sequence.
We designed DNA plasmids containing the mammalian U6 promoter (see Circuit Production for more information on the U6 promoter) that could be used to drive expression of our RNAs in vivo upon transfection into mammalian cells.
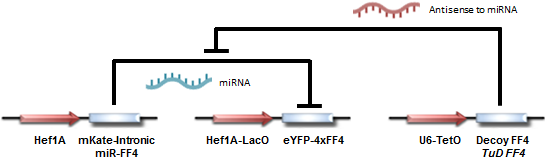
Circuit diagram for testing TuD and Decoy-based actuation.
Our circuit begins with a constitutively (always; from the Hef1A promoter) expressed yellow fluorescent protein (eYFP), with the mRNA transcript containing several binding domains for a particular miRNA (FF4). We then introduce the miRNA (from the Hef1A-LacO:mKate-Intronic miR-FF4 construct) into the system, resulting in repression of yellow protein expression. Finally, upon introducing Decoy and TuD RNA into the system (the U6-TetO:Decoy FF4/TuD FF4 construct), the repression is subsequently relieved, allowing for expression of the yellow fluorescent protein. Note that the miR-FF4 miRNA is produced as a spliced-out intron from mKate, a red fluorescent protein. Thus red fluorescence indicates presence of miR-FF4.
